|
|
Design and synthesis of pleated DNA origami nanotubes with adjustable diameters
Jonathan F. Berengut, Julian C. Berengut, Jonathan P.K. Doye, Domen Prešern, Akiniro Kawamoto, Juanfang Ruan, Madeleine J. Wainwright and Lawrence K. Lee, submitted
Nucleic Acids Res. 47, 11963–11975 (2019)
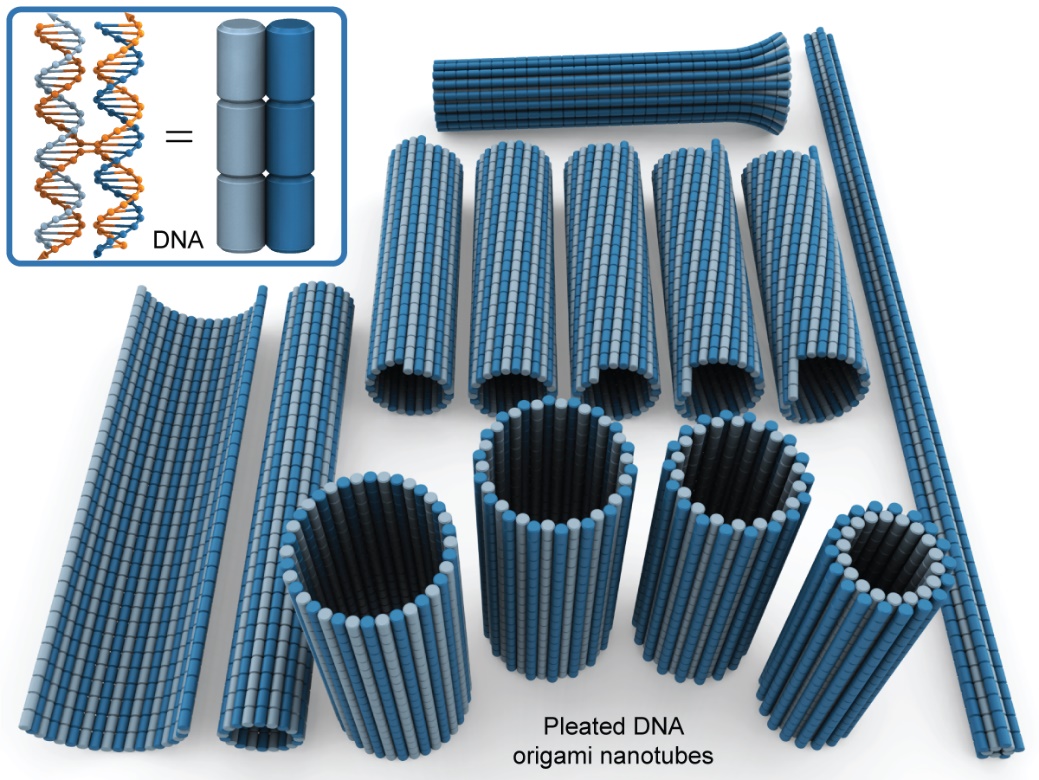
Abstract
DNA origami allows for the synthesis of nanoscale structures and machines with nanometre precision and high yields. Tubular DNA origami nanostructures are particularly useful because their geometry facilitates a variety of applications including nanoparticle encapsulation, the construction of artificial membrane pores and as structural scaffolds that can uniquely spatially arrange nanoparticles in circular, linear and helical arrays. Here we report a system of parametrisation for the design of radially symmetric DNA origami nanotubes with adjustable diameter, length, crossover density, pleat angle and chirality. The system is implemented into a computational algorithm that provides a practical means to navigate the complex geometry of DNA origami nanotube design. We apply this in the design, synthesis and characterisation of novel DNA origami nanotubes. These include structures with pleated walls where the same number of duplexes can form nanotubes with different diameters, and to vary the diameter within the same structure. We also construct nanotubes that can be reconfigured into different chiral shapes. Finally, we explore the effect of strain on the local and global geometry of DNA origami nanotubes and demonstrate how pleated walls can provide a strategy to rigidify nanotubes and to construct closely packed parallel duplexes.The full paper is available from Nucleic Acids Res. and biorXiv.org.