|
|
Coarse-grained modelling of the structural properties of DNA Origami
Antonio Suma, Erik Poppleton, Michael Matthies, Petr Sulc, Flavio Romano, Ard A. Louis, Jonathan P.K. Doye, Christian Micheletti, and Lorenzo Rovigatti
J. Comput. Chem. 40, 2586-2595 (2019)
Abstract
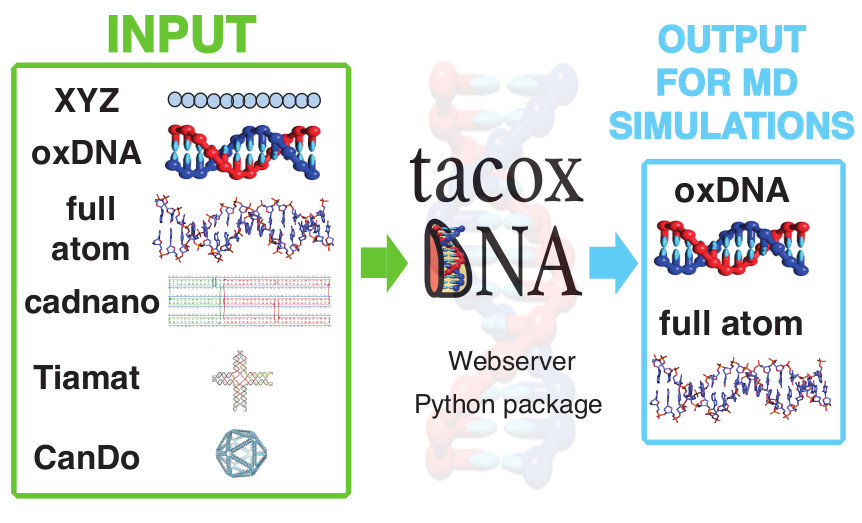
Simulations of nucleic acids at different levels of structural details are increasingly used to complement and interpret experiments in different fields, from biophysics to medicine and materials science. However, the various structural models currently available for DNA and RNA and their accompanying suites of computational tools can be very rarely used in a synergistic fashion. The tacoxDNA webserver and standalone software package presented here are a step towards a long-sought interoperability of nucleic acids models. The webserver offers a simple interface for converting various common input formats of DNA structures and setting up molecular dynamics simulations. Users can, for instance, design DNA rings with different topologies, such as knots, with and without supercoiling, by simply providing an XYZ coordinate file of the DNA centre-line. More complex DNA geometries, as designable in the cadnano, CanDo and Tiamat tools, can also be converted to all-atom or oxDNA representations, which can then be used to run molecular dynamics simulations. Though the latter are currently geared towards the native and LAMMPS oxDNA representations, the open-source package is designed to be further expandable. TacoxDNA is available at http://tacoxdna.sissa.itThe full paper is available from the Journal of Computational Chemistry